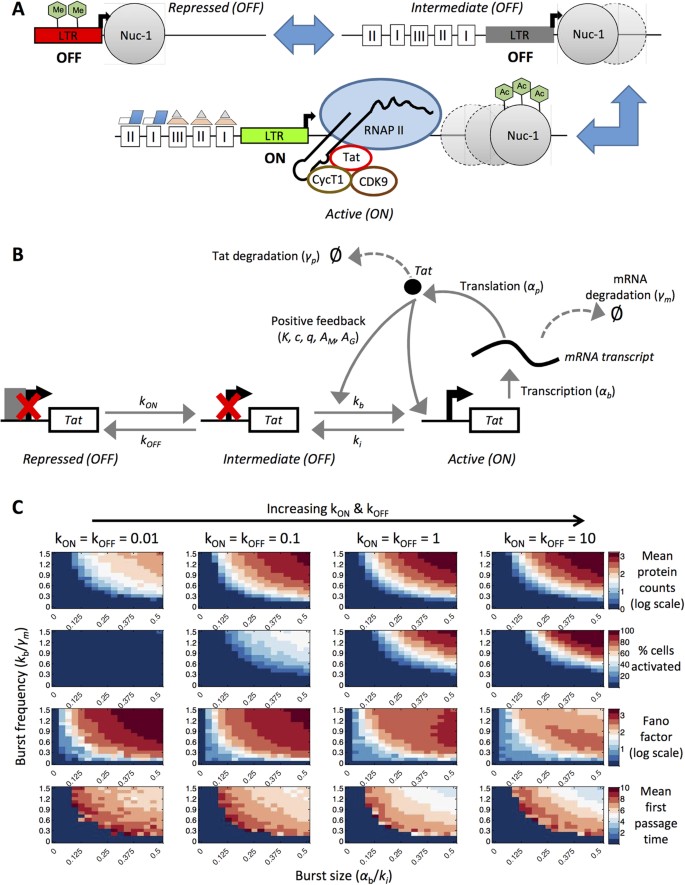
PLOS Computational Biology: Effect of transcription factor resource sharing on gene expression noise

Extrinsic noise prevents the independent tuning of gene expression noise and protein mean abundance in bacteria
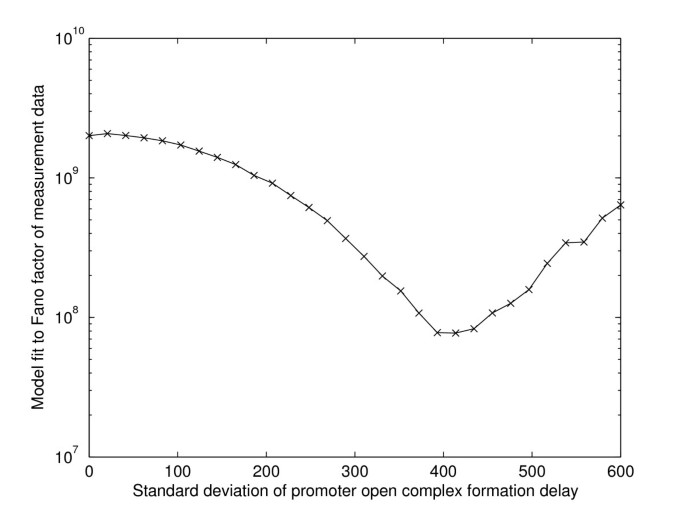
Cell-to-cell diversity in protein levels of a gene driven by a tetracycline inducible promoter | BMC Molecular Biology | Full Text

a–d Variation of Fano factor (calculated from Master equation) with... | Download Scientific Diagram
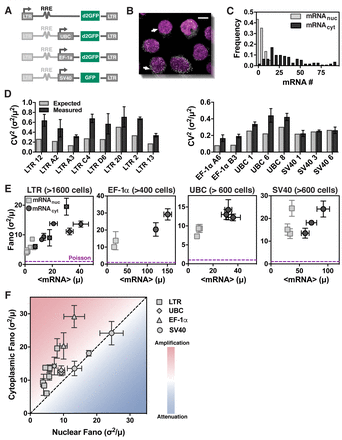
Cytoplasmic import and processing of mRNA amplify transcriptional bursts accounting for the majority of cellular noise | bioRxiv
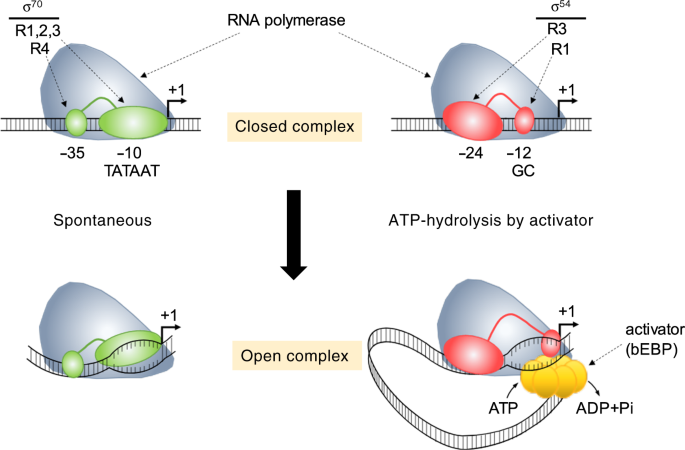
The route to transcription initiation determines the mode of transcriptional bursting in E. coli | Nature Communications

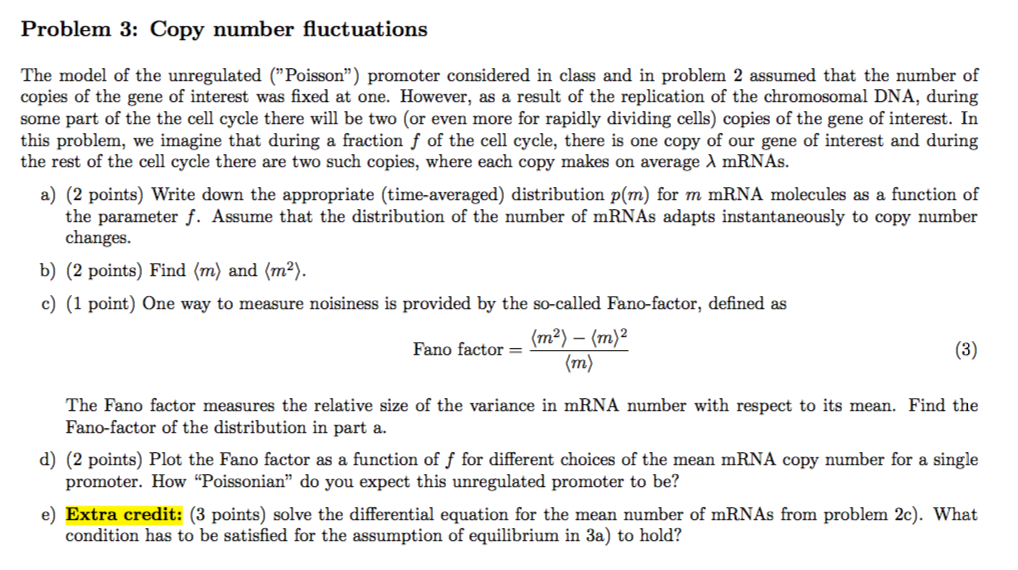
A) Model of transcriptional regulation: The promoter switches between... | Download Scientific Diagram

Promoter-mediated diversification of transcriptional bursting dynamics following gene duplication | PNAS

Reconciling molecular regulatory mechanisms with noise patterns of bacterial metabolic promoters in induced and repressed states | PNAS

mRNA noise of the promoter. (a) Trend of mRNA numbers (Fano factors)... | Download Scientific Diagram

Decoding the grammar of transcriptional regulation from RNA polymerase measurements: models and their applications - IOPscience